Enabling accessible computing, data, and software at scale — for research and education
OneSciencePlace gives institutions, research groups, and educators a single platform for delivering web-based access to apps, data, and computing systems — configured to fit your project, supported by our team.
Jupyter
RStudio
MATLAB
Linux Desktop
Batch / CLI
Publications
Data Sharing
Less to build. Less to maintain. More time for science and education.
OneSciencePlace handles the platform-level work that's typically required to stand up a research site, so your team's effort can stay on research, teaching, and supporting users.
One platform. Four core capabilities.
Compute, Apps, Data, and Publishing — composable to fit your research and educational needs. Start with what your project needs today; add the others as it grows.
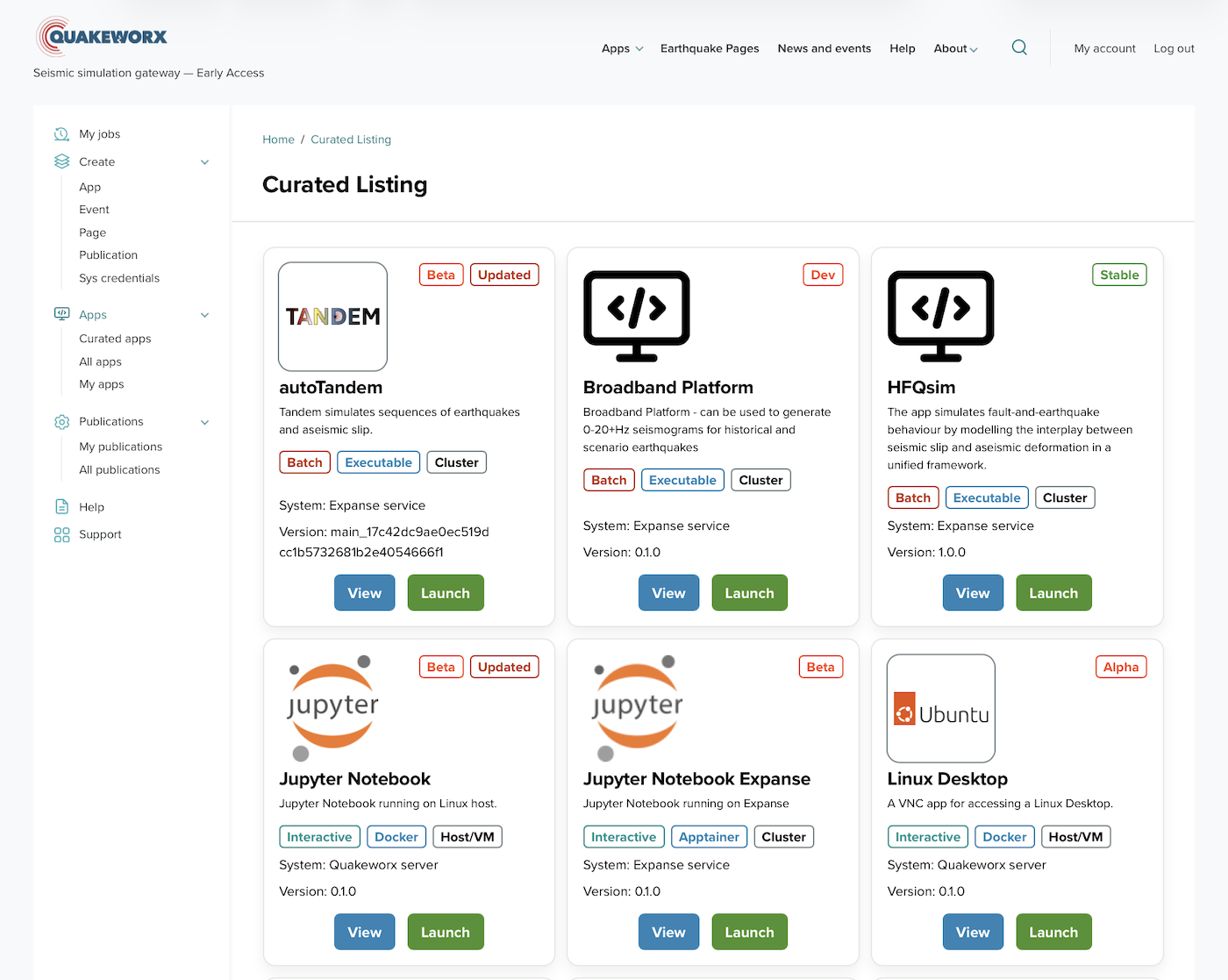
Quakeworx: a browser-based gateway for earthquake simulation
Quakeworx delivers more than twenty domain-specific simulation tools and interactive notebooks to the seismology community through a single browser interface. Built on OneSciencePlace in collaboration with SCEC, USC, UIUC, and Scripps Institution of Oceanography.
Where OSP fits
OneSciencePlace serves a single lab as readily as a multi-institution collaboration. The same platform supports each.
How we deploy with you
OneSciencePlace is available today as a managed deployment. An engagement typically begins with a conversation about your project, and proceeds through five steps.
Technical capabilities
What OneSciencePlace connects to and how, for teams evaluating the technical fit.
Tell us about your project.
We can discuss your needs and how OneSciencePlace might support them.